Genetic drift
| Part of a series on |
| Evolutionary biology |
|---|
 |
Genetic drift, also known as random genetic drift, allelic drift or the Wright effect,[1] is the change in the frequency of an existing gene variant (allele) in a population due to random chance.[2]
Genetic drift may cause gene variants to disappear completely and thereby reduce genetic variation.[3] It can also cause initially rare alleles to become much more frequent and even fixed.
When few copies of an allele exist, the effect of genetic drift is more notable, and when many copies exist, the effect is less notable (due to the law of large numbers). In the middle of the 20th century, vigorous debates occurred over the relative importance of natural selection versus neutral processes, including genetic drift. Ronald Fisher, who explained natural selection using Mendelian genetics,[4] held the view that genetic drift plays at most a minor role in evolution, and this remained the dominant view for several decades. In 1968, population geneticist Motoo Kimura rekindled the debate with his neutral theory of molecular evolution, which claims that most instances where a genetic change spreads across a population (although not necessarily changes in phenotypes) are caused by genetic drift acting on neutral mutations.[5][6] In the 1990s, constructive neutral evolution was proposed which seeks to explain how complex systems emerge through neutral transitions.[7][8]
Analogy with marbles in a jar
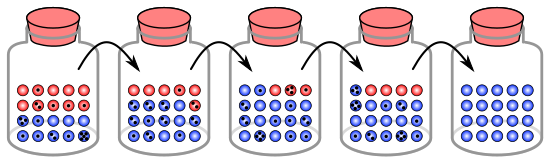
[edit]The process of genetic drift can be illustrated using 20 marbles in a jar to represent 20 organisms in a population.[9] Consider this jar of marbles as the starting population. Half of the marbles in the jar are red and half are blue, with each colour corresponding to a different allele of one gene in the population. In each new generation, the organisms reproduce at random. To represent this reproduction, randomly select a marble from the original jar and deposit a new marble with the same colour into a new jar. This is the "offspring" of the original marble, meaning that the original marble remains in its jar. Repeat this process until 20 new marbles are in the second jar. The second jar will now contain 20 "offspring", or marbles of various colours. Unless the second jar contains exactly 10 red marbles and 10 blue marbles, a random shift has occurred in the allele frequencies.
If this process is repeated a number of times, the numbers of red and blue marbles picked each generation fluctuates. Sometimes, a jar has more red marbles than its "parent" jar and sometimes more blue. This fluctuation is analogous to genetic drift – a change in the population's allele frequency resulting from a random variation in the distribution of alleles from one generation to the next.
In any one generation, no marbles of a particular colour could be chosen, meaning they have no offspring. In this example, if no red marbles are selected, the jar representing the new generation contains only blue offspring. If this happens, the red allele has been lost permanently in the population, while the remaining blue allele has become fixed: all future generations are entirely blue. In small populations, fixation can occur in just a few generations.
Probability and allele frequency
[edit]The mechanisms of genetic drift can be illustrated with a simplified example. Consider a very large colony of bacteria isolated in a drop of solution. The bacteria are genetically identical except for a single gene with two alleles labeled A and B, which are neutral alleles, meaning that they do not affect the bacteria's ability to survive and reproduce; all bacteria in this colony are equally likely to survive and reproduce. Suppose that half the bacteria have allele A and the other half have allele B. Thus, A and B each has an allele frequency of 1/2.
The drop of solution then shrinks until it has only enough food to sustain four bacteria. All other bacteria die without reproducing. Among the four that survive, 16 possible combinations for the A and B alleles exist:
(A-A-A-A), (B-A-A-A), (A-B-A-A), (B-B-A-A),
(A-A-B-A), (B-A-B-A), (A-B-B-A), (B-B-B-A),
(A-A-A-B), (B-A-A-B), (A-B-A-B), (B-B-A-B),
(A-A-B-B), (B-A-B-B), (A-B-B-B), (B-B-B-B).
Since all bacteria in the original solution are equally likely to survive when the solution shrinks, the four survivors are a random sample from the original colony. The probability that each of the four survivors has a given allele is 1/2, and so the probability that any particular allele combination occurs when the solution shrinks is
(The original population size is so large that the sampling effectively happens with replacement). In other words, each of the 16 possible allele combinations is equally likely to occur, with probability 1/16.
Counting the combinations with the same number of A and B gives the following table:
| A | B | Combinations | Probability |
| 4 | 0 | 1 | 1/16 |
| 3 | 1 | 4 | 4/16 |
| 2 | 2 | 6 | 6/16 |
| 1 | 3 | 4 | 4/16 |
| 0 | 4 | 1 | 1/16 |
As shown in the table, the total number of combinations that have the same number of A alleles as of B alleles is six, and the probability of this combination is 6/16. The total number of other combinations is ten, so the probability of unequal number of A and B alleles is 10/16. Thus, although the original colony began with an equal number of A and B alleles, quite possibly, the number of alleles in the remaining population of four members will not be equal. The situation of equal numbers is actually less likely than unequal numbers. In the latter case, genetic drift has occurred because the population's allele frequencies have changed due to random sampling. In this example, the population contracted to just four random survivors, a phenomenon known as a population bottleneck.
The probabilities for the number of copies of allele A (or B) that survive (given in the last column of the above table) can be calculated directly from the binomial distribution, where the "success" probability (probability of a given allele being present) is 1/2 (i.e., the probability that there are k copies of A (or B) alleles in the combination) is given by:
where n=4 is the number of surviving bacteria.
Mathematical models
[edit]Mathematical models of genetic drift can be designed using either branching processes or a diffusion equation describing changes in allele frequency in an idealised population.[10]
Wright–Fisher model
[edit]Consider a gene with two alleles, A or B. In diploidy, populations consisting of N individuals have 2N copies of each gene. An individual can have two copies of the same allele or two different alleles. The frequency of one allele is assigned p and the other q. The Wright–Fisher model (named after Sewall Wright and Ronald Fisher) assumes that generations do not overlap (for example, annual plants have exactly one generation per year) and that each copy of the gene found in the new generation is drawn independently at random from all copies of the gene in the old generation. The formula to calculate the probability of obtaining k copies of an allele that had frequency p in the last generation is then[11][12]
where the symbol "!" signifies the factorial function. This expression can also be formulated using the binomial coefficient,
Moran model
[edit]The Moran model assumes overlapping generations. At each time step, one individual is chosen to reproduce and one individual is chosen to die. So in each timestep, the number of copies of a given allele can go up by one, go down by one, or can stay the same. This means that the transition matrix is tridiagonal, which means that mathematical solutions are easier for the Moran model than for the Wright–Fisher model. On the other hand, computer simulations are usually easier to perform using the Wright–Fisher model, because fewer time steps need to be calculated. In the Moran model, it takes N timesteps to get through one generation, where N is the effective population size. In the Wright–Fisher model, it takes just one.[13]
In practice, the Moran and Wright–Fisher models give qualitatively similar results, but genetic drift runs twice as fast in the Moran model.
Other models of drift
[edit]If the variance in the number of offspring is much greater than that given by the binomial distribution assumed by the Wright–Fisher model, then given the same overall speed of genetic drift (the variance effective population size), genetic drift is a less powerful force compared to selection.[14] Even for the same variance, if higher moments of the offspring number distribution exceed those of the binomial distribution then again the force of genetic drift is substantially weakened.[15]
Random effects other than sampling error
[edit]Random changes in allele frequencies can also be caused by effects other than sampling error, for example random changes in selection pressure.[16]
One important alternative source of stochasticity, perhaps more important than genetic drift, is genetic draft.[17] Genetic draft is the effect on a locus by selection on linked loci. The mathematical properties of genetic draft are different from those of genetic drift.[18] The direction of the random change in allele frequency is autocorrelated across generations.[2]
Drift and fixation
[edit]The Hardy–Weinberg principle states that within sufficiently large populations, the allele frequencies remain constant from one generation to the next unless the equilibrium is disturbed by migration, genetic mutations, or selection.[19]
However, in finite populations, no new alleles are gained from the random sampling of alleles passed to the next generation, but the sampling can cause an existing allele to disappear. Because random sampling can remove, but not replace, an allele, and because random declines or increases in allele frequency influence expected allele distributions for the next generation, genetic drift drives a population towards genetic uniformity over time. When an allele reaches a frequency of 1 (100%) it is said to be "fixed" in the population and when an allele reaches a frequency of 0 (0%) it is lost. Smaller populations achieve fixation faster, whereas in the limit of an infinite population, fixation is not achieved. Once an allele becomes fixed, genetic drift comes to a halt, and the allele frequency cannot change unless a new allele is introduced in the population via mutation or gene flow. Thus even while genetic drift is a random, directionless process, it acts to eliminate genetic variation over time.[20]
Rate of allele frequency change due to drift
[edit]
Assuming genetic drift is the only evolutionary force acting on an allele, after t generations in many replicated populations, starting with allele frequencies of p and q, the variance in allele frequency across those populations is
Time to fixation or loss
[edit]Assuming genetic drift is the only evolutionary force acting on an allele, at any given time the probability that an allele will eventually become fixed in the population is simply its frequency in the population at that time.[22] For example, if the frequency p for allele A is 75% and the frequency q for allele B is 25%, then given unlimited time the probability A will ultimately become fixed in the population is 75% and the probability that B will become fixed is 25%.
The expected number of generations for fixation to occur is proportional to the population size, such that fixation is predicted to occur much more rapidly in smaller populations.[23] Normally the effective population size, which is smaller than the total population, is used to determine these probabilities. The effective population (Ne) takes into account factors such as the level of inbreeding, the stage of the lifecycle in which the population is the smallest, and the fact that some neutral genes are genetically linked to others that are under selection.[14] The effective population size may not be the same for every gene in the same population.[24]
One forward-looking formula used for approximating the expected time before a neutral allele becomes fixed through genetic drift, according to the Wright–Fisher model, is
where T is the number of generations, Ne is the effective population size, and p is the initial frequency for the given allele. The result is the number of generations expected to pass before fixation occurs for a given allele in a population with given size (Ne) and allele frequency (p).[25]
The expected time for the neutral allele to be lost through genetic drift can be calculated as[11]
When a mutation appears only once in a population large enough for the initial frequency to be negligible, the formulas can be simplified to[26]
for average number of generations expected before fixation of a neutral mutation, and
for the average number of generations expected before the loss of a neutral mutation in a population of actual size N.[27]
Time to loss with both drift and mutation
[edit]The formulae above apply to an allele that is already present in a population, and which is subject to neither mutation nor natural selection. If an allele is lost by mutation much more often than it is gained by mutation, then mutation, as well as drift, may influence the time to loss. If the allele prone to mutational loss begins as fixed in the population, and is lost by mutation at rate m per replication, then the expected time in generations until its loss in a haploid population is given by
where is Euler's constant.[28] The first approximation represents the waiting time until the first mutant destined for loss, with loss then occurring relatively rapidly by genetic drift, taking time 1/m ≫ Ne. The second approximation represents the time needed for deterministic loss by mutation accumulation. In both cases, the time to fixation is dominated by mutation via the term 1/m, and is less affected by the effective population size.
Versus natural selection
[edit]In natural populations, genetic drift and natural selection do not act in isolation; both phenomena are always at play, together with mutation and migration. Neutral evolution is the product of both mutation and drift, not of drift alone. Similarly, even when selection overwhelms genetic drift, it can only act on variation that mutation provides.
While natural selection has a direction, guiding evolution towards heritable adaptations to the current environment, genetic drift has no direction and is guided only by the mathematics of chance.[29] As a result, drift acts upon the genotypic frequencies within a population without regard to their phenotypic effects. In contrast, selection favors the spread of alleles whose phenotypic effects increase survival and/or reproduction of their carriers, lowers the frequencies of alleles that cause unfavorable traits, and ignores those that are neutral.[30]
The law of large numbers predicts that when the absolute number of copies of the allele is small (e.g., in small populations), the magnitude of drift on allele frequencies per generation is larger. The magnitude of drift is large enough to overwhelm selection at any allele frequency when the selection coefficient is less than 1 divided by the effective population size. Non-adaptive evolution resulting from the product of mutation and genetic drift is therefore considered to be a consequential mechanism of evolutionary change primarily within small, isolated populations.[31] The mathematics of genetic drift depend on the effective population size, but it is not clear how this is related to the actual number of individuals in a population.[17] Genetic linkage to other genes that are under selection can reduce the effective population size experienced by a neutral allele. With a higher recombination rate, linkage decreases and with it this local effect on effective population size.[32][33] This effect is visible in molecular data as a correlation between local recombination rate and genetic diversity,[34] and negative correlation between gene density and diversity at noncoding DNA regions.[35] Stochasticity associated with linkage to other genes that are under selection is not the same as sampling error, and is sometimes known as genetic draft in order to distinguish it from genetic drift.[17]
Low allele frequency makes alleles more vulnerable to being eliminated by random chance, even overriding the influence of natural selection. For example, while disadvantageous mutations are usually eliminated quickly within the population, new advantageous mutations are almost as vulnerable to loss through genetic drift as are neutral mutations. Not until the allele frequency for the advantageous mutation reaches a certain threshold will genetic drift have no effect.[30]
Population bottleneck
[edit]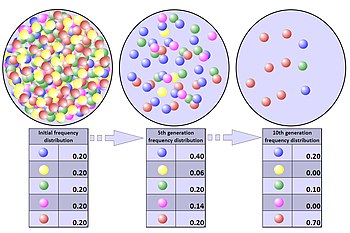
A population bottleneck is when a population contracts to a significantly smaller size over a short period of time due to some random environmental event. In a true population bottleneck, the odds for survival of any member of the population are purely random, and are not improved by any particular inherent genetic advantage. The bottleneck can result in radical changes in allele frequencies, completely independent of selection.[36]
The impact of a population bottleneck can be sustained, even when the bottleneck is caused by a one-time event such as a natural catastrophe. An interesting example of a bottleneck causing unusual genetic distribution is the relatively high proportion of individuals with total rod cell color blindness (achromatopsia) on Pingelap atoll in Micronesia.[37] After a bottleneck, inbreeding increases. This increases the damage done by recessive deleterious mutations, in a process known as inbreeding depression. The worst of these mutations are selected against, leading to the loss of other alleles that are genetically linked to them, in a process of background selection.[2] For recessive harmful mutations, this selection can be enhanced as a consequence of the bottleneck, due to genetic purging. This leads to a further loss of genetic diversity. In addition, a sustained reduction in population size increases the likelihood of further allele fluctuations from drift in generations to come.
A population's genetic variation can be greatly reduced by a bottleneck, and even beneficial adaptations may be permanently eliminated.[38] The loss of variation leaves the surviving population vulnerable to any new selection pressures such as disease, climatic change or shift in the available food source, because adapting in response to environmental changes requires sufficient genetic variation in the population for natural selection to take place.[39][40]
There have been many known cases of population bottleneck in the recent past. Prior to the arrival of Europeans, North American prairies were habitat for millions of greater prairie chickens. In Illinois alone, their numbers plummeted from about 100 million birds in 1900 to about 50 birds in the 1990s. The declines in population resulted from hunting and habitat destruction, but a consequence has been a loss of most of the species' genetic diversity. DNA analysis comparing birds from the mid century to birds in the 1990s documents a steep decline in the genetic variation in just the latter few decades. Currently the greater prairie chicken is experiencing low reproductive success.[41]
However, the genetic loss caused by bottleneck and genetic drift can increase fitness, as in Ehrlichia.[42]
Over-hunting also caused a severe population bottleneck in the northern elephant seal in the 19th century. Their resulting decline in genetic variation can be deduced by comparing it to that of the southern elephant seal, which were not so aggressively hunted.[43]
Founder effect
[edit]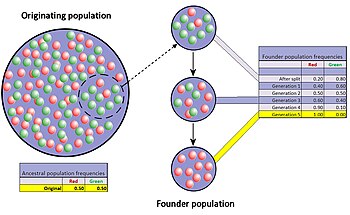
The founder effect is a special case of a population bottleneck, occurring when a small group in a population splinters off from the original population and forms a new one. The random sample of alleles in the just formed new colony is expected to grossly misrepresent the original population in at least some respects.[44] It is even possible that the number of alleles for some genes in the original population is larger than the number of gene copies in the founders, making complete representation impossible. When a newly formed colony is small, its founders can strongly affect the population's genetic make-up far into the future.
A well-documented example is found in the Amish migration to Pennsylvania in 1744. Two members of the new colony shared the recessive allele for Ellis–Van Creveld syndrome. Members of the colony and their descendants tend to be religious isolates and remain relatively insular. As a result of many generations of inbreeding, Ellis–Van Creveld syndrome is now much more prevalent among the Amish than in the general population.[30][45]
The difference in gene frequencies between the original population and colony may also trigger the two groups to diverge significantly over the course of many generations. As the difference, or genetic distance, increases, the two separated populations may become distinct, both genetically and phenetically, although not only genetic drift but also natural selection, gene flow, and mutation contribute to this divergence. This potential for relatively rapid changes in the colony's gene frequency led most scientists to consider the founder effect (and by extension, genetic drift) a significant driving force in the evolution of new species. Sewall Wright was the first to attach this significance to random drift and small, newly isolated populations with his shifting balance theory of speciation.[46] Following after Wright, Ernst Mayr created many persuasive models to show that the decline in genetic variation and small population size following the founder effect were critically important for new species to develop.[47] However, there is much less support for this view today since the hypothesis has been tested repeatedly through experimental research and the results have been equivocal at best.[48]
History
[edit]The role of random chance in evolution was first outlined by Arend L. Hagedoorn and Anna Cornelia Hagedoorn-Vorstheuvel La Brand in 1921.[49] They highlighted that random survival plays a key role in the loss of variation from populations. Fisher (1922) responded to this with the first, albeit marginally incorrect, mathematical treatment of the "Hagedoorn effect".[50] Notably, he expected that many natural populations were too large (an N ~10,000) for the effects of drift to be substantial and thought drift would have an insignificant effect on the evolutionary process. The corrected mathematical treatment and term "genetic drift" was later coined by a founder of population genetics, Sewall Wright. His first use of the term "drift" was in 1929,[51] though at the time he was using it in the sense of a directed process of change, or natural selection. Random drift by means of sampling error came to be known as the "Sewall–Wright effect," though he was never entirely comfortable to see his name given to it. Wright referred to all changes in allele frequency as either "steady drift" (e.g., selection) or "random drift" (e.g., sampling error).[52] "Drift" came to be adopted as a technical term in the stochastic sense exclusively.[53] Today it is usually defined still more narrowly, in terms of sampling error,[54] although this narrow definition is not universal.[55][56] Wright wrote that the "restriction of "random drift" or even "drift" to only one component, the effects of accidents of sampling, tends to lead to confusion".[52] Sewall Wright considered the process of random genetic drift by means of sampling error equivalent to that by means of inbreeding, but later work has shown them to be distinct.[57]
In the early days of the modern evolutionary synthesis, scientists were beginning to blend the new science of population genetics with Charles Darwin's theory of natural selection. Within this framework, Wright focused on the effects of inbreeding on small relatively isolated populations. He introduced the concept of an adaptive landscape in which phenomena such as cross breeding and genetic drift in small populations could push them away from adaptive peaks, which in turn allow natural selection to push them towards new adaptive peaks.[58] Wright thought smaller populations were more suited for natural selection because "inbreeding was sufficiently intense to create new interaction systems through random drift but not intense enough to cause random nonadaptive fixation of genes".[59]
Wright's views on the role of genetic drift in the evolutionary scheme were controversial almost from the very beginning. One of the most vociferous and influential critics was colleague Ronald Fisher. Fisher conceded genetic drift played some role in evolution, but an insignificant one. Fisher has been accused of misunderstanding Wright's views because in his criticisms Fisher seemed to argue Wright had rejected selection almost entirely. To Fisher, viewing the process of evolution as a long, steady, adaptive progression was the only way to explain the ever-increasing complexity from simpler forms. But the debates have continued between the "gradualists" and those who lean more toward the Wright model of evolution where selection and drift together play an important role.[60]
In 1968, Motoo Kimura rekindled the debate with his neutral theory of molecular evolution, which claims that most of the genetic changes are caused by genetic drift acting on neutral mutations.[5][6]
The role of genetic drift by means of sampling error in evolution has been criticized by John H. Gillespie[61] and William B. Provine, who argue that selection on linked sites is a more important stochastic force.
See also
[edit]- Peripatric speciation
- Antigenic drift
- Coalescent theory
- Constructive neutral evolution
- Gene pool
- Meiotic drive
Notes and references
[edit]- ^ Gould SJ (2002). "Chapter 7, section "Synthesis as Hardening"". The Structure of Evolutionary Theory.
- ^ a b c Masel J (October 2011). "Genetic drift". Current Biology. 21 (20). Cell Press: R837-8. doi:10.1016/j.cub.2011.08.007. PMID 22032182.
- ^ Star B, Spencer HG (May 2013). "Effects of genetic drift and gene flow on the selective maintenance of genetic variation". Genetics. 194 (1): 235–44. doi:10.1534/genetics.113.149781. PMC 3632471. PMID 23457235.
- ^ Miller 2000, p. 54
- ^ a b Kimura M (February 1968). "Evolutionary rate at the molecular level". Nature. 217 (5129). Nature Publishing Group: 624–6. Bibcode:1968Natur.217..624K. doi:10.1038/217624a0. PMID 5637732. S2CID 4161261.
- ^ a b Futuyma 1998, p. 320
- ^ Stoltzfus A (1999). "On the Possibility of Constructive Neutral Evolution". Journal of Molecular Evolution. 49 (2): 169–181. Bibcode:1999JMolE..49..169S. doi:10.1007/PL00006540. ISSN 0022-2844. PMID 10441669. S2CID 1743092. Archived from the original on 30 July 2022. Retrieved 20 January 2022.
- ^ Muñoz-Gómez SA, Bilolikar G, Wideman JG, Geiler-Samerotte K (April 2021). "Constructive Neutral Evolution 20 Years Later". Journal of Molecular Evolution. 89 (3): 172–182. Bibcode:2021JMolE..89..172M. doi:10.1007/s00239-021-09996-y. PMC 7982386. PMID 33604782.
- ^ "Sampling Error and Evolution". Understanding Evolution. University of California, Berkeley. Archived from the original on 8 December 2015. Retrieved 1 December 2015.
- ^ Wahl LM (August 2011). "Fixation when N and s vary: classic approaches give elegant new results". Genetics. 188 (4). Genetics Society of America: 783–5. doi:10.1534/genetics.111.131748. PMC 3176088. PMID 21828279.
- ^ a b Hartl & Clark 2007, p. 112
- ^ Tian 2008, p. 11
- ^ Moran PA (1958). "Random processes in genetics". Mathematical Proceedings of the Cambridge Philosophical Society. 54 (1): 60–71. Bibcode:1958PCPS...54...60M. doi:10.1017/S0305004100033193. S2CID 85823386.
- ^ a b Charlesworth B (March 2009). "Fundamental concepts in genetics: effective population size and patterns of molecular evolution and variation". Nature Reviews. Genetics. 10 (3). Nature Publishing Group: 195–205. doi:10.1038/nrg2526. PMID 19204717. S2CID 205484393.
- ^ Der R, Epstein CL, Plotkin JB (September 2011). "Generalized population models and the nature of genetic drift". Theoretical Population Biology. 80 (2). Elsevier: 80–99. doi:10.1016/j.tpb.2011.06.004. PMID 21718713.
- ^ Li & Graur 1991, p. 28
- ^ a b c Gillespie JH (November 2001). "Is the population size of a species relevant to its evolution?". Evolution; International Journal of Organic Evolution. 55 (11). John Wiley & Sons for the Society for the Study of Evolution: 2161–9. doi:10.1111/j.0014-3820.2001.tb00732.x. PMID 11794777. S2CID 221735887.
- ^ Neher RA, Shraiman BI (August 2011). "Genetic draft and quasi-neutrality in large facultatively sexual populations". Genetics. 188 (4). Genetics Society of America: 975–96. arXiv:1108.1635. doi:10.1534/genetics.111.128876. PMC 3176096. PMID 21625002.
- ^ Ewens 2004
- ^ Li & Graur 1991, p. 29
- ^ Barton et al. 2007, p. 417
- ^ Futuyma 1998, p. 300
- ^ Otto SP, Whitlock MC (June 1997). "The probability of fixation in populations of changing size" (PDF). Genetics. 146 (2). Genetics Society of America: 723–33. doi:10.1093/genetics/146.2.723. PMC 1208011. PMID 9178020. Archived (PDF) from the original on 19 March 2015.
- ^ Cutter AD, Choi JY (August 2010). "Natural selection shapes nucleotide polymorphism across the genome of the nematode Caenorhabditis briggsae". Genome Research. 20 (8). Cold Spring Harbor Laboratory Press: 1103–11. doi:10.1101/gr.104331.109. PMC 2909573. PMID 20508143.
- ^ Hedrick 2005, p. 315
- ^ Li & Graur 1991, p. 33
- ^ Kimura & Ohta 1971
- ^ Masel J, King OD, Maughan H (January 2007). "The loss of adaptive plasticity during long periods of environmental stasis". The American Naturalist. 169 (1). University of Chicago Press on behalf of the American Society of Naturalists: 38–46. doi:10.1086/510212. PMC 1766558. PMID 17206583.
- ^ "Natural Selection: How Evolution Works". Actionbioscience. Washington, D.C.: American Institute of Biological Sciences. Archived from the original on 6 January 2010. Retrieved 24 November 2009. An interview with Douglas J. Futuyma. See answer to question: Is natural selection the only mechanism of evolution?
- ^ a b c Cavalli-Sforza, Menozzi & Piazza 1996
- ^ Zimmer 2001
- ^ Golding 1994, p. 46
- ^ Charlesworth B, Morgan MT, Charlesworth D (August 1993). "The effect of deleterious mutations on neutral molecular variation" (PDF). Genetics. 134 (4). Genetics Society of America: 1289–303. doi:10.1093/genetics/134.4.1289. PMC 1205596. PMID 8375663. Archived (PDF) from the original on 12 March 2020. Retrieved 9 December 2015.
- ^ Presgraves DC (September 2005). "Recombination enhances protein adaptation in Drosophila melanogaster". Current Biology. 15 (18). Cell Press: 1651–6. doi:10.1016/j.cub.2005.07.065. PMID 16169487. S2CID 15120927.
- ^ Nordborg M, Hu TT, Ishino Y, Jhaveri J, Toomajian C, Zheng H, Bakker E, Calabrese P, Gladstone J, Goyal R, Jakobsson M, Kim S, Morozov Y, Padhukasahasram B, Plagnol V, Rosenberg NA, Shah C, Wall JD, Wang J, Zhao K, Kalbfleisch T, Schulz V, Kreitman M, Bergelson J (July 2005). "The pattern of polymorphism in Arabidopsis thaliana". PLOS Biology. 3 (7). Public Library of Science: e196. doi:10.1371/journal.pbio.0030196. PMC 1135296. PMID 15907155.

- ^ Robinson R, ed. (2003). "Population Bottleneck". Genetics. Vol. 3. New York: Macmillan Reference USA. ISBN 0-02-865609-1. LCCN 2002003560. OCLC 614996575. Retrieved 14 December 2015.
- ^ Hussels IE, Morton NE (May 1972). "Pingelap and Mokil Atolls: achromatopsia". American Journal of Human Genetics. 24 (3): 304–309. PMC 1762260. PMID 4555088.
- ^ Futuyma 1998, pp. 303–304
- ^ O'Corry-Crowe G (March 2008). "Climate change and the molecular ecology of Arctic marine mammals". Ecological Applications. 18 (2 Suppl). Ecological Society of America: S56-76. doi:10.1890/06-0795.1. PMID 18494363.
- ^ Cornuet JM, Luikart G (December 1996). "Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data". Genetics. 144 (4). Genetics Society of America: 2001–14. doi:10.1093/genetics/144.4.2001. PMC 1207747. PMID 8978083.
- ^ Sadava et al. 2008, chpts. 1, 21–33, 52–57
- ^ Dale C, Moran NA (August 2006). "Molecular interactions between bacterial symbionts and their hosts". Cell. 126 (3): 453–65. doi:10.1016/j.cell.2006.07.014. PMID 16901780. S2CID 15985536.
- ^ "Bottlenecks and founder effects". Understanding Evolution. University of California, Berkeley. Archived from the original on 4 December 2015. Retrieved 14 December 2015.
- ^ Campbell 1996, p. 423
- ^ "Genetic Drift and the Founder Effect". Evolution Library (Web resource). Evolution. Boston, MA: WGBH Educational Foundation; Clear Blue Sky Productions, Inc. 2001. OCLC 48165595. Archived from the original on 14 March 2009. Retrieved 7 April 2009.
- ^ Wolf, Brodie & Wade 2000
- ^ Hey, Fitch & Ayala 2005
- ^ Howard & Berlocher 1998
- ^ Hagedoorn AL, Hagedoorn-Vorstheuvel La Brand AC (1921). The Relative Value of the Processes Causing Evolution. The Hague: Martinus Nijhoff. Archived from the original on 28 March 2019. Retrieved 28 March 2019.
- ^ Fisher RA (1922). "On the Dominance Ratio". Proceedings of the Royal Society of Edinburgh. 42: 321–341. doi:10.1017/s0370164600023993.
- ^ Wright S (November–December 1929). "The evolution of dominance". The American Naturalist. 63 (689). Chicago, IL: University of Chicago Press on behalf of the American Society of Naturalists: 556–561. doi:10.1086/280290. ISSN 0003-0147. JSTOR 2456825. S2CID 85301374.
- ^ a b Wright S (1955). "Classification of the factors of evolution". Cold Spring Harbor Symposia on Quantitative Biology. 20. Cold Spring Harbor Laboratory Press: 16–24D. doi:10.1101/SQB.1955.020.01.004. PMID 13433551. Symposium: "Population Genetics: The Nature and Causes of Genetic Variability in Populations".
- ^ Stevenson 1991
- ^ Freeman & Herron 2007
- ^ Masel J (August 2012). "Rethinking Hardy-Weinberg and genetic drift in undergraduate biology". BioEssays. 34 (8). John Wiley & Sons: 701–710. doi:10.1002/bies.201100178. PMID 22576789. S2CID 28513167.
- ^ Lynch 2007
- ^ Crow JF (March 2010). "Wright and Fisher on inbreeding and random drift". Genetics. 184 (3). Genetics Society of America: 609–611. doi:10.1534/genetics.109.110023. PMC 2845331. PMID 20332416.
- ^ Larson 2004, pp. 221–243
- ^ Stevenson 1991: Quote attributed to William B. Provine in The Origins of Theoretical Population Genetics (1971), p. 162; Chicago: University of Chicago Press.
- ^ Avers 1989
- ^ Gillespie JH (June 2000). "Genetic drift in an infinite population. The pseudohitchhiking model". Genetics. 155 (2). Genetics Society of America: 909–919. doi:10.1093/genetics/155.2.909. PMC 1461093. PMID 10835409.
Bibliography
[edit]- Avers CJ (1989). Process and Pattern in Evolution. New York: Oxford University Press. ISBN 0-19-505275-7. LCCN 88005368. OCLC 17677554.
- Barton NH, Briggs DE, Eisen JA, Goldstein DB, Patel NH (2007). Evolution. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press. ISBN 978-0-87969-684-9. LCCN 2007010767. OCLC 86090399.
- Campbell NA (1996). Biology. Benjamin/Cummings Series in the Life Sciences (4th ed.). Menlo Park, CA: Benjamin/Cummings Pub. Co. ISBN 0-8053-1940-9. LCCN 95045572. OCLC 33333455.
- Cavalli-Sforza LL, Menozzi P, Piazza A (1996). The History and Geography of Human Genes (Abridged paperback ed.). Princeton, N.J.: Princeton University Press. ISBN 0-691-02905-9. OCLC 35527063.
- Ewens WJ (2004). Mathematical Population Genetics I. Theoretical Introduction. Interdisciplinary Applied Mathematics. Vol. 27 (2nd ed.). New York: Springer-Verlag. ISBN 0-387-20191-2. LCCN 2003065728. OCLC 53231891.
- Freeman S, Herron JC (2007). Evolutionary Analysis (4th ed.). Upper Saddle River, NJ: Pearson Prentice Hall. ISBN 978-0-13-227584-2. LCCN 2006034384. OCLC 73502978.
- Futuyma D (1998). Evolutionary Biology (3rd ed.). Sunderland, MA: Sinauer Associates. ISBN 0-87893-189-9. LCCN 97037947. OCLC 37560100.
- Golding B, ed. (1994). Non-Neutral Evolution: Theories and Molecular Data. New York: Chapman & Hall. ISBN 0-412-05391-8. LCCN 93047006. OCLC 29638235. "Papers from a workshop sponsored by the Canadian Institute for Advanced Research."
- Hartl DL, Clark AG (2007). Principles of Population Genetics (4th ed.). Sunderland, MA: Sinauer Associates. ISBN 978-0-87893-308-2. LCCN 2006036153. OCLC 75087956.
- Hedrick PW (2005). Genetics of Populations (3rd ed.). Boston, MA: Jones and Bartlett Publishers. ISBN 0-7637-4772-6. LCCN 2004056666. OCLC 56194719.
- Hey J, Fitch WM, Ayala FJ, eds. (2005). Systematics and the Origin of Species: On Ernst Mayr's 100th Anniversary. Washington, D.C.: National Academies Press. ISBN 978-0-309-09536-5. LCCN 2005017917. OCLC 70745851.
- Howard DJ, Berlocher SH, eds. (1998). Endless Forms: Species and Speciation. New York: Oxford University Press. ISBN 978-0-19-510901-6. LCCN 97031461. OCLC 37545522.
- Kimura M, Ohta T (1971). Theoretical Aspects of Population Genetics. Monographs in Population Biology. Vol. 4. Princeton, NJ: Princeton University Press. pp. 1–219. ISBN 0-691-08096-8. LCCN 75155963. OCLC 299867647. PMID 5162676.
- Larson EJ (2004). Evolution: The Remarkable History of a Scientific Theory. Modern Library Chronicles. Vol. 17. New York: Modern Library. ISBN 0-679-64288-9. LCCN 2003064888. OCLC 53483597.
- Li WH, Graur D (1991). Fundamentals of Molecular Evolution. Sunderland, MA: Sinauer Associates. ISBN 0-87893-452-9. LCCN 90043581. OCLC 22113526.
- Lynch M (2007). The Origins of Genome Architecture. Sunderland, MA: Sinauer Associates. ISBN 978-0-87893-484-3. LCCN 2007000012. OCLC 77574049.
- Miller G (2000). The Mating Mind: How Sexual Choice Shaped the Evolution of Human Nature. New York: Doubleday. ISBN 0-385-49516-1. LCCN 00022673. OCLC 43648482.
- Sadava D, Heller HC, Orians GH, Purves WK, Hillis DM (2008). Life: The Science of Biology. Vol. II: Evolution, Diversity and Ecology (8th ed.). Sunderland, MA; Gordonsville, VA: Sinauer Associates; W. H. Freeman and Company. ISBN 978-0-7167-7674-1. LCCN 2006031320. OCLC 71632224.
- Stevenson JC (1991). Dictionary of Concepts in Physical Anthropology. Reference Sources for the Social Sciences and Humanities. Vol. 10. Westport, CT: Greenwood Press. ISBN 0-313-24756-0. LCCN 90022815. OCLC 22732327.
- Tian JP (2008). Evolution Algebras and their Applications. Lecture Notes in Mathematics. Vol. 1921. Berlin; New York: Springer. doi:10.1007/978-3-540-74284-5. ISBN 978-3-540-74283-8. LCCN 2007933498. OCLC 173807298. Zbl 1136.17001.
- Wolf JB, Brodie ED, Wade MJ, eds. (2000). Epistasis and the Evolutionary Process. Oxford, UK; New York: Oxford University Press. ISBN 0-19-512806-0. LCCN 99046515. OCLC 42603105.
- Zimmer C (2001). Evolution: The Triumph of an Idea. Introduction by Stephen Jay Gould; foreword by Richard Hutton (1st ed.). New York: HarperCollins. ISBN 0-06-019906-7. LCCN 2001024077. OCLC 46359440.
External links
[edit]- Sheehy, Bob. "Population genetics simulation program". Radford, VA: Radford University. Retrieved 21 December 2015.
- Grimes, Bill. "Genetic Drift Simulation". Tucson, Arizona: The University of Arizona. Retrieved 25 August 2016.










![{\displaystyle {\bar {T}}_{\text{lost}}\approx {\begin{cases}{\dfrac {1}{m}},{\text{ if }}mN_{e}\ll 1\\[8pt]{\dfrac {\ln {(mN_{e})}+\gamma }{m}}{\text{ if }}mN_{e}\gg 1\end{cases}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/0dca9b7dc746a7b2a8d8a63770ac53782d1639e3)
